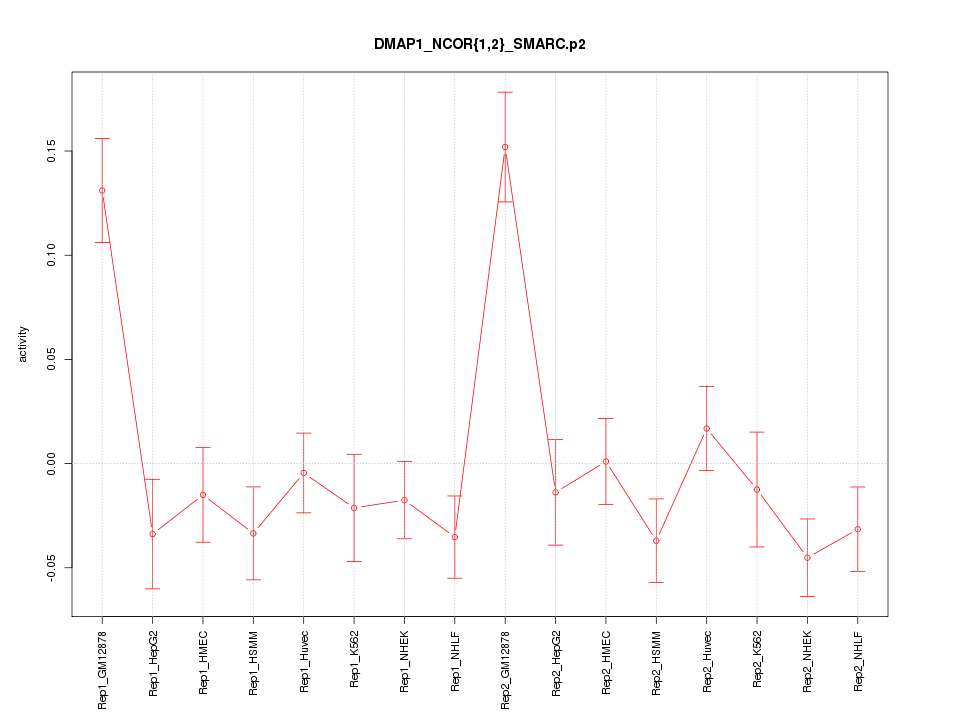
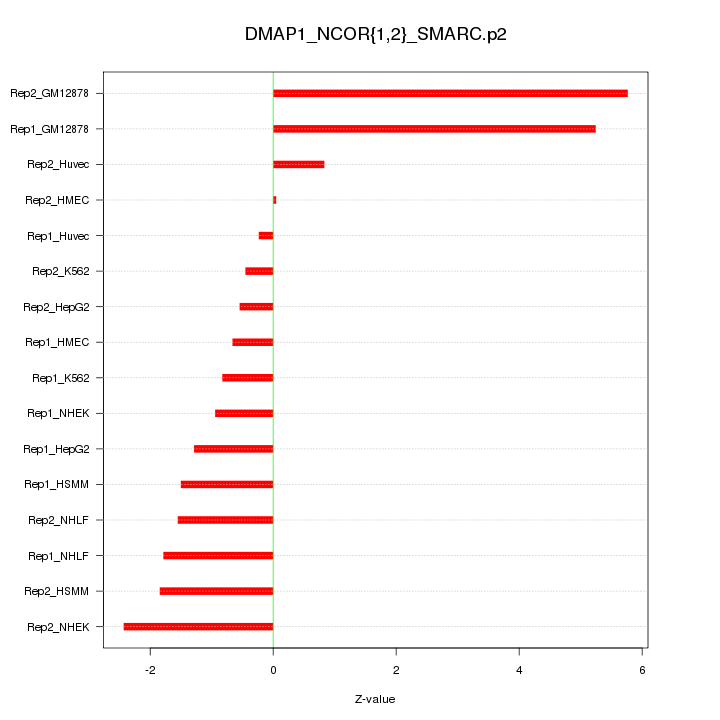
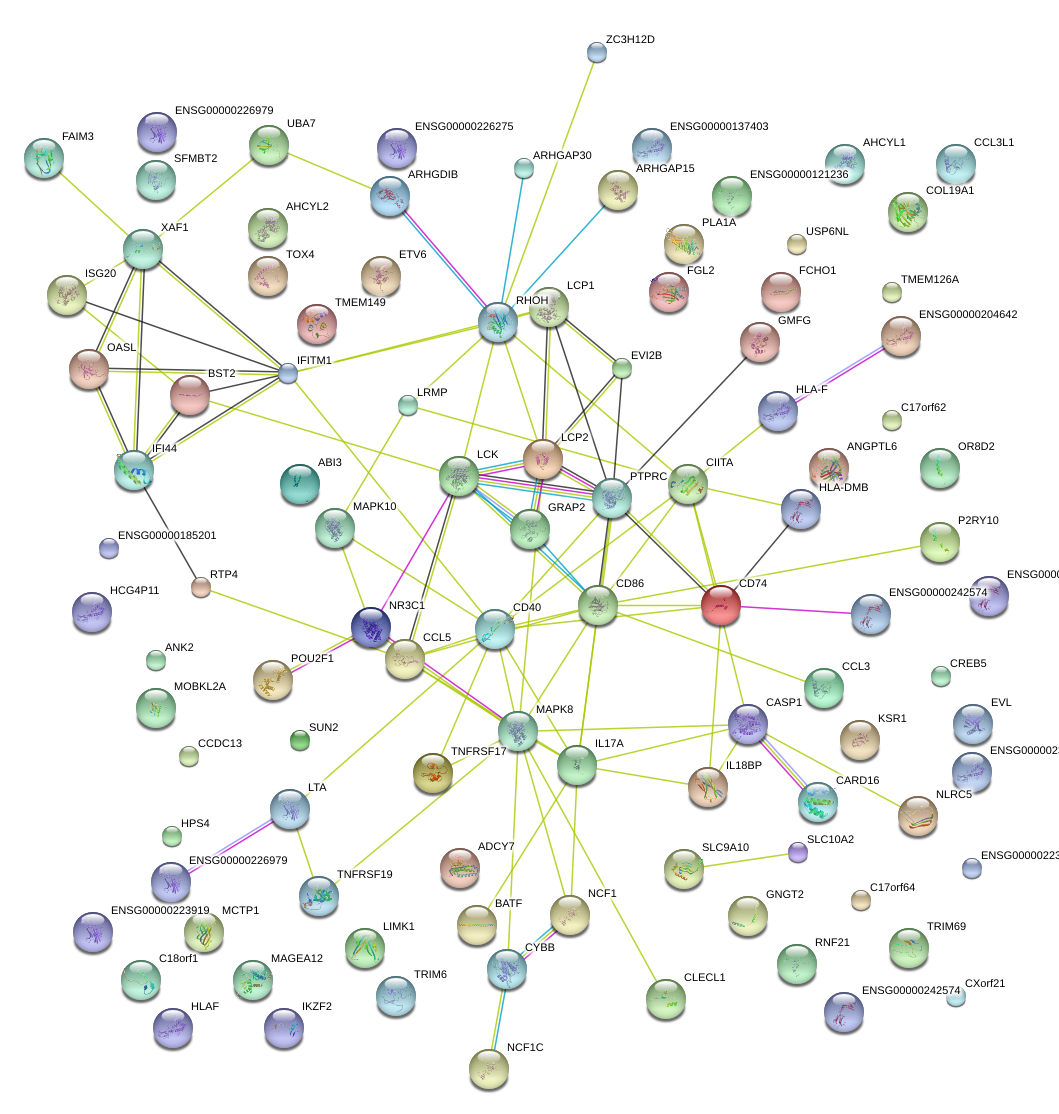
|
chr15_+_42815851
|
6.717
|
NM_080745
NM_182985
|
TRIM69
|
tripartite motif containing 69
|
|
chr3_+_120799411
|
6.193
|
NM_015900
|
PLA1A
|
phospholipase A1 member A
|
|
chrX_+_78087484
|
5.597
|
NM_014499
NM_198333
|
P2RY10
|
purinergic receptor P2Y, G-protein coupled, 10
|
|
chr5_-_138753443
|
5.397
|
NM_016459
|
MZB1
|
marginal zone B and B1 cell-specific protein
|
|
chr10_-_11614279
|
5.309
|
NM_001080491
|
USP6NL
|
USP6 N-terminal like
|
|
chr16_+_48865521
|
4.869
|
|
ADCY7
|
adenylate cyclase 7
|
|
chr3_+_123279439
|
4.829
|
NM_006889
|
CD86
|
CD86 molecule
|
|
chrX_-_30505804
|
4.763
|
NM_025159
|
CXorf21
|
chromosome X open reading frame 21
|
|
chr16_+_10878539
|
4.514
|
NM_000246
|
CIITA
|
class II, major histocompatibility complex, transactivator
|
|
chr7_+_73826244
|
4.490
|
NM_000265
|
NCF1
NCF1B
NCF1C
|
neutrophil cytosolic factor 1
neutrophil cytosolic factor 1B pseudogene
neutrophil cytosolic factor 1C pseudogene
|
|
chr13_+_23042722
|
4.482
|
NM_148957
|
TNFRSF19
|
tumor necrosis factor receptor superfamily, member 19
|
|
chr2_+_143603439
|
3.966
|
|
ARHGAP15
|
Rho GTPase activating protein 15
|
|
chr14_+_99601498
|
3.869
|
NM_016337
|
EVL
|
Enah/Vasp-like
|
|
chr14_+_99601517
|
3.869
|
|
EVL
|
Enah/Vasp-like
|
|
chr2_+_143603368
|
3.820
|
NM_018460
|
ARHGAP15
|
Rho GTPase activating protein 15
|
|
chr19_+_17377502
|
3.772
|
|
LOC100507535
|
hypothetical LOC100507535
|
|
chr14_+_75058518
|
3.732
|
NM_006399
|
BATF
|
basic leucine zipper transcription factor, ATF-like
|
|
chr12_-_15005761
|
3.698
|
NM_001175
|
ARHGDIB
|
Rho GDP dissociation inhibitor (GDI) beta
|
|
chr14_+_75058591
|
3.688
|
|
BATF
|
basic leucine zipper transcription factor, ATF-like
|
|
chr5_-_169657312
|
3.650
|
|
LCP2
|
lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa)
|
|
chr22_-_25205376
|
3.647
|
NM_152841
|
HPS4
|
Hermansky-Pudlak syndrome 4
|
|
chr1_+_165564904
|
3.641
|
NM_001198783
|
POU2F1
|
POU class 2 homeobox 1
|
|
chr1_+_165565006
|
3.587
|
|
POU2F1
|
POU class 2 homeobox 1
|
|
chr1_-_205161821
|
3.526
|
NM_001193338
NM_001142473
NM_005449
|
FAIM3
|
Fas apoptotic inhibitory molecule 3
|
|
chr17_-_78000890
|
3.516
|
NM_001193655
|
C17orf62
|
chromosome 17 open reading frame 62
|
|
chr5_-_169657192
|
3.281
|
|
LCP2
|
lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa)
|
|
chr17_+_55854636
|
3.208
|
NM_181707
|
C17orf64
|
chromosome 17 open reading frame 64
|
|
chr6_-_33016561
|
3.200
|
NM_002118
|
HLA-DMB
|
major histocompatibility complex, class II, DM beta
|
|
chr5_-_169657395
|
3.046
|
NM_005565
|
LCP2
|
lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa)
|
|
chr17_+_44643067
|
3.040
|
|
ABI3
|
ABI family, member 3
|
|
chr19_-_10074332
|
3.012
|
NM_031917
|
ANGPTL6
|
angiopoietin-like 6
|
|
chr20_+_44180355
|
2.912
|
|
CD40
|
CD40 molecule, TNF receptor superfamily member 5
|
|
chr20_+_44180296
|
2.908
|
NM_001250
NM_152854
|
CD40
|
CD40 molecule, TNF receptor superfamily member 5
|
|
chr5_-_149772528
|
2.847
|
|
|
|
|
chr5_-_169657360
|
2.830
|
|
LCP2
|
lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa)
|
|
chr5_-_149772469
|
2.749
|
NM_001025159
NM_004355
NM_001025158
|
CD74
|
CD74 molecule, major histocompatibility complex, class II invariant chain
|
|
chr17_-_44642934
|
2.645
|
NM_001198754
|
GNGT2
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 2
|
|
chr19_-_2036227
|
2.611
|
|
MOBKL2A
|
MOB1, Mps One Binder kinase activator-like 2A (yeast)
|
|
chr1_-_159306327
|
2.588
|
|
ARHGAP30
|
Rho GTPase activating protein 30
|
|
chr4_+_39874921
|
2.582
|
NM_004310
|
RHOH
|
ras homolog gene family, member H
|
|
chr19_+_17723285
|
2.579
|
NM_001161359
NM_015122
|
FCHO1
|
FCH domain only 1
|
|
chr17_+_6600080
|
2.576
|
|
XAF1
|
XIAP associated factor 1
|
|
chr6_+_70633168
|
2.503
|
NM_001858
|
COL19A1
|
collagen, type XIX, alpha 1
|
|
chr6_+_31647854
|
2.452
|
NM_001159740
|
LTA
|
lymphotoxin alpha (TNF superfamily, member 1)
|
|
chr17_+_22823151
|
2.448
|
NM_014238
|
KSR1
|
kinase suppressor of ras 1
|
|
chr1_+_196874839
|
2.446
|
|
PTPRC
|
protein tyrosine phosphatase, receptor type, C
|
|
chr1_+_32489426
|
2.413
|
NM_005356
|
LCK
|
lymphocyte-specific protein tyrosine kinase
|
|
chr11_+_5602782
|
2.386
|
NM_021616
|
TRIM34
|
tripartite motif containing 34
|
|
chr19_-_44518465
|
2.374
|
|
GMFG
|
glia maturation factor, gamma
|
|
chr6_+_52159143
|
2.342
|
NM_002190
|
IL17A
|
interleukin 17A
|
|
chr7_+_73145481
|
2.333
|
|
LIMK1
|
LIM domain kinase 1
|
|
chr19_-_44518516
|
2.325
|
NM_004877
|
GMFG
|
glia maturation factor, gamma
|
|
chr15_+_86983038
|
2.324
|
NM_002201
|
ISG20
|
interferon stimulated exonuclease gene 20kDa
|
|
chrX_+_37524190
|
2.323
|
NM_000397
|
CYBB
|
cytochrome b-245, beta polypeptide
|
|
chr11_+_303852
|
2.311
|
NM_003641
|
IFITM1
|
interferon induced transmembrane protein 1 (9-27)
|
|
chr12_+_25096472
|
2.200
|
NM_006152
|
LRMP
|
lymphoid-restricted membrane protein
|
|
chr1_-_159306369
|
2.147
|
NM_001025598
NM_181720
|
ARHGAP30
|
Rho GTPase activating protein 30
|
|
chr18_+_13601664
|
2.092
|
NM_001003674
NM_001003675
|
C18orf1
|
chromosome 18 open reading frame 1
|
|
chr5_-_142795269
|
2.030
|
NM_001018074
NM_001018075
NM_001018077
|
NR3C1
|
nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor)
|
|
chr12_-_119961143
|
2.023
|
NM_003733
NM_198213
|
OASL
|
2'-5'-oligoadenylate synthetase-like
|
|
chr19_-_40925151
|
2.010
|
NM_024660
|
TMEM149
|
transmembrane protein 149
|
|
chr7_+_128795178
|
1.983
|
NM_001130723
|
AHCYL2
|
adenosylhomocysteinase-like 2
|
|
chr12_-_9777126
|
1.976
|
NM_172004
|
CLECL1
|
C-type lectin-like 1
|
|
chrX_+_37524250
|
1.951
|
|
CYBB
|
cytochrome b-245, beta polypeptide
|
|
chr7_+_128795280
|
1.917
|
|
AHCYL2
|
adenosylhomocysteinase-like 2
|
|
chr6_+_29799182
|
1.914
|
|
HLA-F
|
major histocompatibility complex, class I, F
|
|
chr17_-_31231421
|
1.914
|
NM_002985
|
CCL5
|
chemokine (C-C motif) ligand 5
|
|
chr7_-_76667045
|
1.896
|
NM_006682
|
FGL2
|
fibrinogen-like 2
|
|
chr19_-_17377383
|
1.883
|
NM_004335
|
BST2
|
bone marrow stromal cell antigen 2
|
|
chr13_-_102517196
|
1.849
|
NM_000452
|
SLC10A2
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 2
|
|
chr22_+_38627031
|
1.845
|
NM_004810
|
GRAP2
|
GRB2-related adaptor protein 2
|
|
chr3_-_49826221
|
1.828
|
|
UBA7
|
ubiquitin-like modifier activating enzyme 7
|
|
chr9_+_126460511
|
1.825
|
|
|
|
|
chr7_+_28418668
|
1.818
|
NM_182898
|
CREB5
|
cAMP responsive element binding protein 5
|
|
chr2_-_213723298
|
1.806
|
NM_016260
|
IKZF2
|
IKAROS family zinc finger 2 (Helios)
|
|
chr5_-_149772563
|
1.778
|
|
CD74
|
CD74 molecule, major histocompatibility complex, class II invariant chain
|
|
chr3_-_113495763
|
1.778
|
NM_183061
|
SLC9A10
|
solute carrier family 9, member 10
|
|
chr17_-_26665213
|
1.773
|
|
EVI2B
|
ecotropic viral integration site 2B
|
|
chr3_+_188568861
|
1.764
|
NM_022147
|
RTP4
|
receptor (chemosensory) transporter protein 4
|
|
chr1_+_78888064
|
1.691
|
NM_006417
|
IFI44
|
interferon-induced protein 44
|
|
chr12_+_11694172
|
1.667
|
|
ETV6
|
ets variant 6
|
|
chr13_-_45654327
|
1.661
|
NM_002298
|
LCP1
|
lymphocyte cytosolic protein 1 (L-plastin)
|
|
chr1_+_196874759
|
1.641
|
NM_002838
NM_080921
NM_080923
|
PTPRC
|
protein tyrosine phosphatase, receptor type, C
|
|
chr5_-_94442941
|
1.588
|
|
MCTP1
|
multiple C2 domains, transmembrane 1
|
|
chr16_+_11966464
|
1.558
|
NM_001192
|
TNFRSF17
|
tumor necrosis factor receptor superfamily, member 17
|
|
chr17_-_31441390
|
1.531
|
NM_002983
|
CCL3
|
chemokine (C-C motif) ligand 3
|
|
chr16_+_55580910
|
1.512
|
NM_032206
|
NLRC5
|
NLR family, CARD domain containing 5
|
|
chr11_-_104421240
|
1.473
|
NM_001017534
NM_052889
|
CARD16
CASP1
|
caspase recruitment domain family, member 16
caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase)
|
|
chr6_-_149847718
|
1.464
|
|
ZC3H12D
|
zinc finger CCCH-type containing 12D
|
|
chr3_-_42789738
|
1.451
|
NM_144719
|
CCDC13
|
coiled-coil domain containing 13
|
|
chr11_-_104411065
|
1.433
|
NM_001223
NM_033292
NM_033293
NM_033294
NM_033295
|
CASP1
|
caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase)
|
|
chr4_+_113958652
|
1.431
|
NM_001127493
|
ANK2
|
ankyrin 2, neuronal
|
|
chr10_-_7491204
|
1.408
|
NM_001018039
|
SFMBT2
|
Scm-like with four mbt domains 2
|
|
chr11_+_71387572
|
1.406
|
NM_001039659
NM_001039660
NM_173042
|
IL18BP
|
interleukin 18 binding protein
|
|
chr22_+_38627129
|
1.395
|
|
GRAP2
|
GRB2-related adaptor protein 2
|
|
chr4_-_87500201
|
1.389
|
|
MAPK10
|
mitogen-activated protein kinase 10
|
|
chr22_-_37480898
|
1.382
|
|
SUN2
|
Sad1 and UNC84 domain containing 2
|
|
chr11_+_71388310
|
1.372
|
NM_001145057
|
IL18BP
|
interleukin 18 binding protein
|
|
chr1_-_153098811
|
1.362
|
NM_170782
|
KCNN3
|
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3
|
|
chr4_-_153820535
|
1.343
|
NM_152680
|
TMEM154
|
transmembrane protein 154
|
|
chr17_+_7402788
|
1.320
|
|
TNFSF13
|
tumor necrosis factor (ligand) superfamily, member 13
|
|
chr7_+_119700924
|
1.315
|
NM_012281
|
KCND2
|
potassium voltage-gated channel, Shal-related subfamily, member 2
|
|
chr12_-_47604930
|
1.310
|
NM_001143781
|
FKBP11
|
FK506 binding protein 11, 19 kDa
|
|
chr17_+_6599879
|
1.301
|
NM_017523
NM_199139
|
XAF1
|
XIAP associated factor 1
|
|
chr1_-_149004916
|
1.292
|
NM_004079
|
CTSS
|
cathepsin S
|
|
chr18_+_71051713
|
1.282
|
NM_005786
|
TSHZ1
|
teashirt zinc finger homeobox 1
|
|
chr10_-_1769669
|
1.277
|
NM_018702
|
ADARB2
|
adenosine deaminase, RNA-specific, B2
|
|
chr6_-_138231006
|
1.271
|
|
|
|
|
chr1_-_9052253
|
1.257
|
NM_001135585
NM_003039
|
SLC2A5
|
solute carrier family 2 (facilitated glucose/fructose transporter), member 5
|
|
chr2_+_218641982
|
1.254
|
NM_198483
|
RUFY4
|
RUN and FYVE domain containing 4
|
|
chr17_-_37518213
|
1.253
|
NM_024119
|
DHX58
|
DEXH (Asp-Glu-X-His) box polypeptide 58
|
|
chr1_-_9052170
|
1.233
|
|
SLC2A5
|
solute carrier family 2 (facilitated glucose/fructose transporter), member 5
|
|
chr6_-_149847809
|
1.219
|
NM_207360
|
ZC3H12D
|
zinc finger CCCH-type containing 12D
|
|
chr3_+_190831909
|
1.212
|
NM_001114978
NM_001114979
NM_003722
|
TP63
|
tumor protein p63
|
|
chr11_+_128068982
|
1.211
|
NM_002017
|
FLI1
|
Friend leukemia virus integration 1
|
|
chr17_+_44642587
|
1.208
|
NM_001135186
NM_016428
|
ABI3
|
ABI family, member 3
|
|
chr11_+_5667494
|
1.194
|
NM_006074
|
TRIM22
|
tripartite motif containing 22
|
|
chr12_+_6431437
|
1.191
|
NM_018009
|
TAPBPL
|
TAP binding protein-like
|
|
chr17_-_26665144
|
1.180
|
|
EVI2B
|
ecotropic viral integration site 2B
|
|
chr16_-_28425632
|
1.170
|
NM_145659
|
IL27
|
interleukin 27
|
|
chr4_-_38482805
|
1.161
|
NM_003263
|
TLR1
|
toll-like receptor 1
|
|
chr1_+_84539876
|
1.154
|
NM_001134663
|
SAMD13
|
sterile alpha motif domain containing 13
|
|
chr2_-_37752731
|
1.144
|
NM_006449
|
CDC42EP3
|
CDC42 effector protein (Rho GTPase binding) 3
|
|
chr1_-_181889070
|
1.143
|
NM_203454
|
APOBEC4
|
apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 4 (putative)
|
|
chr5_-_88155360
|
1.138
|
NM_001193348
NM_001193349
|
MEF2C
|
myocyte enhancer factor 2C
|
|
chr16_-_49272741
|
1.135
|
NM_001144972
NM_153337
NM_182854
|
SNX20
|
sorting nexin 20
|
|
chr1_-_149004879
|
1.130
|
|
CTSS
|
cathepsin S
|
|
chr1_+_159943385
|
1.128
|
NM_001184866
NM_001184867
NM_001184870
NM_001184871
NM_001184872
NM_001184873
NM_032738
|
FCRLA
|
Fc receptor-like A
|
|
chr12_+_91620749
|
1.121
|
NM_001037671
NM_001178097
|
C12orf74
|
chromosome 12 open reading frame 74
|
|
chrX_-_131401319
|
1.119
|
NM_018388
NM_133486
|
MBNL3
|
muscleblind-like 3 (Drosophila)
|
|
chr1_+_157441824
|
1.116
|
NM_001122951
|
DARC
|
Duffy blood group, chemokine receptor
|
|
chr3_-_114176401
|
1.094
|
NM_138806
NM_138939
NM_138940
NM_170780
|
CD200R1
|
CD200 receptor 1
|
|
chr1_-_149586298
|
1.093
|
|
RFX5
|
regulatory factor X, 5 (influences HLA class II expression)
|
|
chr12_-_623056
|
1.083
|
|
|
|
|
chr19_+_40895669
|
1.074
|
NM_014383
|
ZBTB32
|
zinc finger and BTB domain containing 32
|
|
chr19_+_10057972
|
1.065
|
|
C19orf66
|
chromosome 19 open reading frame 66
|
|
chr12_+_119562737
|
1.063
|
NM_001033677
|
CABP1
|
calcium binding protein 1
|
|
chr1_+_938709
|
1.048
|
NM_005101
|
ISG15
|
ISG15 ubiquitin-like modifier
|
|
chr20_-_51633042
|
1.046
|
NM_006526
|
ZNF217
|
zinc finger protein 217
|
|
chr13_+_23742846
|
1.031
|
|
SPATA13
|
spermatogenesis associated 13
|
|
chr10_+_114125955
|
1.024
|
|
ACSL5
|
acyl-CoA synthetase long-chain family member 5
|
|
chr16_-_88358946
|
1.021
|
|
FANCA
|
Fanconi anemia, complementation group A
|
|
chr12_-_51887318
|
1.020
|
|
ITGB7
|
integrin, beta 7
|
|
chr12_-_47605478
|
1.016
|
NM_001143782
NM_016594
|
FKBP11
|
FK506 binding protein 11, 19 kDa
|
|
chr14_+_23700261
|
1.004
|
NM_006084
|
IRF9
|
interferon regulatory factor 9
|
|
chr11_+_71388619
|
1.002
|
NM_001145055
|
IL18BP
|
interleukin 18 binding protein
|
|
chr19_-_55104875
|
0.995
|
|
NUP62
|
nucleoporin 62kDa
|
|
chr12_+_25096755
|
0.993
|
|
LRMP
|
lymphoid-restricted membrane protein
|
|
chr1_-_25164061
|
0.991
|
NM_001031680
|
RUNX3
|
runt-related transcription factor 3
|
|
chr5_-_137502975
|
0.991
|
NM_003551
|
NME5
|
non-metastatic cells 5, protein expressed in (nucleoside-diphosphate kinase)
|
|
chr11_+_63060848
|
0.986
|
NM_004585
|
RARRES3
|
retinoic acid receptor responder (tazarotene induced) 3
|
|
chr4_+_38740246
|
0.983
|
NM_015990
NM_199039
|
KLHL5
|
kelch-like 5 (Drosophila)
|
|
chr1_-_149004852
|
0.970
|
|
CTSS
|
cathepsin S
|
|
chr7_+_150013716
|
0.954
|
NM_015660
|
GIMAP2
|
GTPase, IMAP family member 2
|
|
chr10_+_35466715
|
0.954
|
NM_183011
NM_183012
|
CREM
|
cAMP responsive element modulator
|
|
chr10_+_91082218
|
0.944
|
NM_001031683
|
IFIT3
|
interferon-induced protein with tetratricopeptide repeats 3
|
|
chr1_+_67404756
|
0.934
|
NM_144701
|
IL23R
|
interleukin 23 receptor
|
|
chr7_+_106293112
|
0.929
|
NM_002649
|
PIK3CG
|
phosphoinositide-3-kinase, catalytic, gamma polypeptide
|
|
chr17_-_26665194
|
0.927
|
|
EVI2B
|
ecotropic viral integration site 2B
|
|
chr12_-_51887266
|
0.919
|
NM_000889
|
ITGB7
|
integrin, beta 7
|
|
chr16_+_29726962
|
0.911
|
|
MAZ
|
MYC-associated zinc finger protein (purine-binding transcription factor)
|
|
chr6_+_29799095
|
0.910
|
NM_001098478
NM_001098479
NM_018950
|
HLA-F
|
major histocompatibility complex, class I, F
|
|
chr1_+_100957883
|
0.908
|
NM_001078
NM_080682
|
VCAM1
|
vascular cell adhesion molecule 1
|
|
chr16_+_29726931
|
0.896
|
|
MAZ
|
MYC-associated zinc finger protein (purine-binding transcription factor)
|
|
chr8_-_102872372
|
0.881
|
|
NCALD
|
neurocalcin delta
|
|
chr3_-_58175437
|
0.878
|
NM_004944
|
DNASE1L3
|
deoxyribonuclease I-like 3
|
|
chr14_-_22358752
|
0.877
|
NM_001126106
|
SLC7A7
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 7
|
|
chr9_+_127550298
|
0.853
|
NM_001134778
|
PBX3
|
pre-B-cell leukemia homeobox 3
|
|
chr1_+_196874867
|
0.852
|
|
PTPRC
|
protein tyrosine phosphatase, receptor type, C
|
|
chr1_-_149586341
|
0.849
|
NM_000449
NM_001025603
|
RFX5
|
regulatory factor X, 5 (influences HLA class II expression)
|
|
chr8_-_102872354
|
0.846
|
|
NCALD
|
neurocalcin delta
|
|
chr15_-_42790805
|
0.844
|
|
|
|
|
chr1_-_159099090
|
0.844
|
NM_001166663
NM_001166664
NM_016382
|
CD244
|
CD244 molecule, natural killer cell receptor 2B4
|
|
chr2_+_182030510
|
0.840
|
|
ITGA4
|
integrin, alpha 4 (antigen CD49D, alpha 4 subunit of VLA-4 receptor)
|
|
chr9_-_5294579
|
0.837
|
NM_005059
NM_134441
|
RLN2
|
relaxin 2
|
|
chr17_-_39094787
|
0.831
|
NM_001040002
|
MEOX1
|
mesenchyme homeobox 1
|
|
chr8_-_145132582
|
0.812
|
NM_032789
|
PARP10
|
poly (ADP-ribose) polymerase family, member 10
|
|
chr17_-_26665220
|
0.805
|
NM_006495
|
EVI2B
|
ecotropic viral integration site 2B
|
|
chr3_-_49370682
|
0.804
|
|
GPX1
|
glutathione peroxidase 1
|
|
chr16_+_27346079
|
0.798
|
NM_021798
|
IL21R
|
interleukin 21 receptor
|
|
chr19_+_10058023
|
0.797
|
|
C19orf66
|
chromosome 19 open reading frame 66
|
|
chr5_-_94443300
|
0.782
|
NM_001002796
|
MCTP1
|
multiple C2 domains, transmembrane 1
|
|
chr8_-_145132551
|
0.781
|
|
PARP10
|
poly (ADP-ribose) polymerase family, member 10
|
|
chr19_+_40511909
|
0.774
|
NM_001185099
NM_001185100
NM_001185101
NM_001771
|
CD22
|
CD22 molecule
|
|
chr5_-_176671437
|
0.769
|
|
MXD3
|
MAX dimerization protein 3
|
|
chr3_+_99733432
|
0.769
|
NM_005290
|
GPR15
|
G protein-coupled receptor 15
|
|
chr13_-_45654296
|
0.765
|
|
LCP1
|
lymphocyte cytosolic protein 1 (L-plastin)
|
|
chr12_+_49922592
|
0.765
|
|
DAZAP2
|
DAZ associated protein 2
|
|
chr10_+_5812616
|
0.762
|
|
C10orf18
|
chromosome 10 open reading frame 18
|
|
chr1_-_246869181
|
0.761
|
NM_001001827
|
OR2T35
|
olfactory receptor, family 2, subfamily T, member 35
|
|
chrX_-_54841397
|
0.757
|
NM_198510
|
ITIH5L
|
inter-alpha (globulin) inhibitor H5-like
|
|
chr4_-_186970366
|
0.753
|
NM_003603
NM_001145670
|
SORBS2
|
sorbin and SH3 domain containing 2
|
|
chr3_-_49826360
|
0.750
|
NM_003335
|
UBA7
|
ubiquitin-like modifier activating enzyme 7
|
|
chr3_+_186563529
|
0.748
|
NM_004721
|
MAP3K13
|
mitogen-activated protein kinase kinase kinase 13
|
|
chr11_-_88436463
|
0.744
|
NM_000842
|
GRM5
|
glutamate receptor, metabotropic 5
|
|
chr11_-_64327197
|
0.741
|
NM_004579
|
MAP4K2
|
mitogen-activated protein kinase kinase kinase kinase 2
|
|
chr17_-_44043278
|
0.736
|
|
HOXB7
|
homeobox B7
|
|
chr6_+_154449334
|
0.732
|
NM_001145287
|
OPRM1
|
opioid receptor, mu 1
|
|
chr2_-_215711313
|
0.722
|
NM_173076
|
ABCA12
|
ATP-binding cassette, sub-family A (ABC1), member 12
|
|
chr14_+_69307914
|
0.722
|
|
SRSF5
|
serine/arginine-rich splicing factor 5
|